|
|
|
| Home » Transcription » Research |
| Transcription |
Current Research Interests
The laboratory of transcription is engaged in understanding the mechanism, physiology, and inhibition of bacterial Rho-dependent termination. A wide range of techniques from biophysics (spectroscopy, thermodynamics, fast kinetics, etc.), biochemistry (protein purification, chemical and enzymatic foot-printing of protein and nucleic acids, cross-linking, etc.), molecular biology (recombinant DNA techniques, site-directed mutagenesis, super-resolution microscopy), bacterial genetics and genomics are used in the laboratory to solve these intellectually challenging problems.
Projects:
- Mechanism of transcription termination by transcription termination factor Rho.
- Mechanism of Rho-NusG interaction in vivo and in vitro.
- Physiological roles of Rho-dependent terminations.
- Super-resolution microscopy of the transcription machinery.
- Fast-kinetics approach to study the transcription termination processes.
- Isolation of mycobacteriophage-derived proteins with antimicrobial activities.
- Design of antimicrobial peptides from bacteriophage proteins.
- Characterization of Rho protein of B.Subtillis
Collaborators::
- Prof Markus Wahl, Freie Universität Berlin, Germany (Cryo-EM).
- Prof. Udaydittya Sen, SINP, Kolkata (Crystallography).
- Prof. Agnieszka Szalewska-Palas, Uniwersytet Gdanski, Poland.
- Prof. Jayanta Mukhopadhyay.
Research Highlights |
|
Peptides designed from a bacteriophage capsid protein function as synthetic transcription repressors (JBC, 2023).
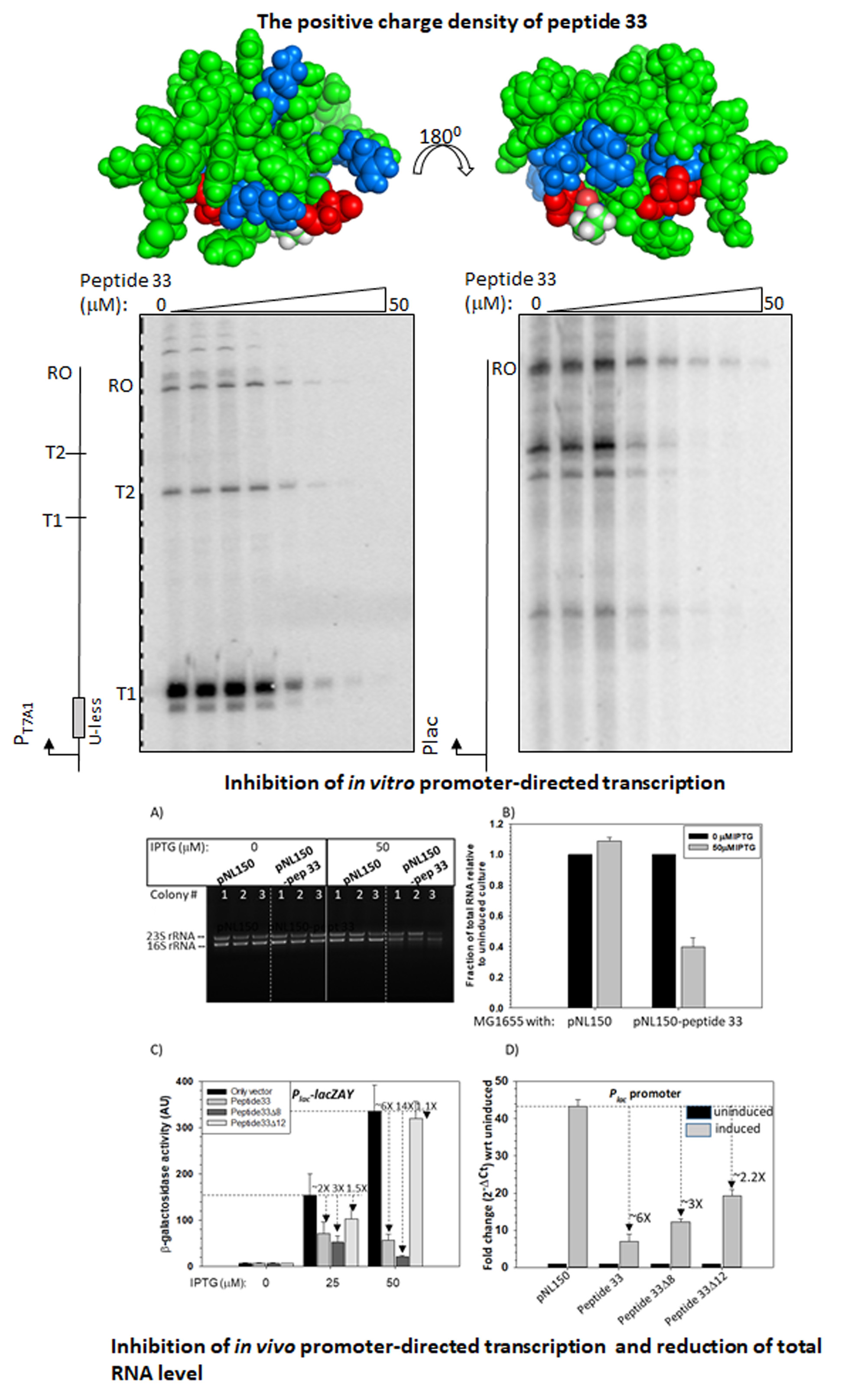
The mycobacteriophages encode unique proteins that are potent therapeutic agents. We screened several clones with mycobactericidal properties from a genomic library of mycobacteriophages. Here we report the properties of one such clone coding the gene product, Gp49, of the phage Che12. Gp49 is a 16 kD dimeric protein having an HTH motif at its C-terminal and is highly conserved among mycobacteriophages and likely to be part of phage DNA replication machinery. Alphafold predicts it to be an -helical protein. However, its CD spectrum showed it to be predominantly -sheeted. It is a high-affinity heparin-binding protein that has similarities with the macrophage protein Azurocidin. Its -sheeted apo-structure gets transformed into -helix upon binding to heparin. It binds to linear dsDNA as well as ssDNA and RNA cooperatively in a sequence non-specific manner. This DNA binding property enables it to inhibit both in vitro and in vivo transcription. The c-terminal HTH motif is responsible for binding to both heparin and nucleic acids. Its in vivo localization on DNA could cause displacements of many DNA-binding proteins from the bacterial chromosome. We surmised that the bactericidal activity of Gp49 arises from its non-specific DNA binding leading to the inhibition of many host-DNA-dependent processes. Its heparin-binding ability could have therapeutic/diagnostic usages in bacterial sepsis treatment.
|
|
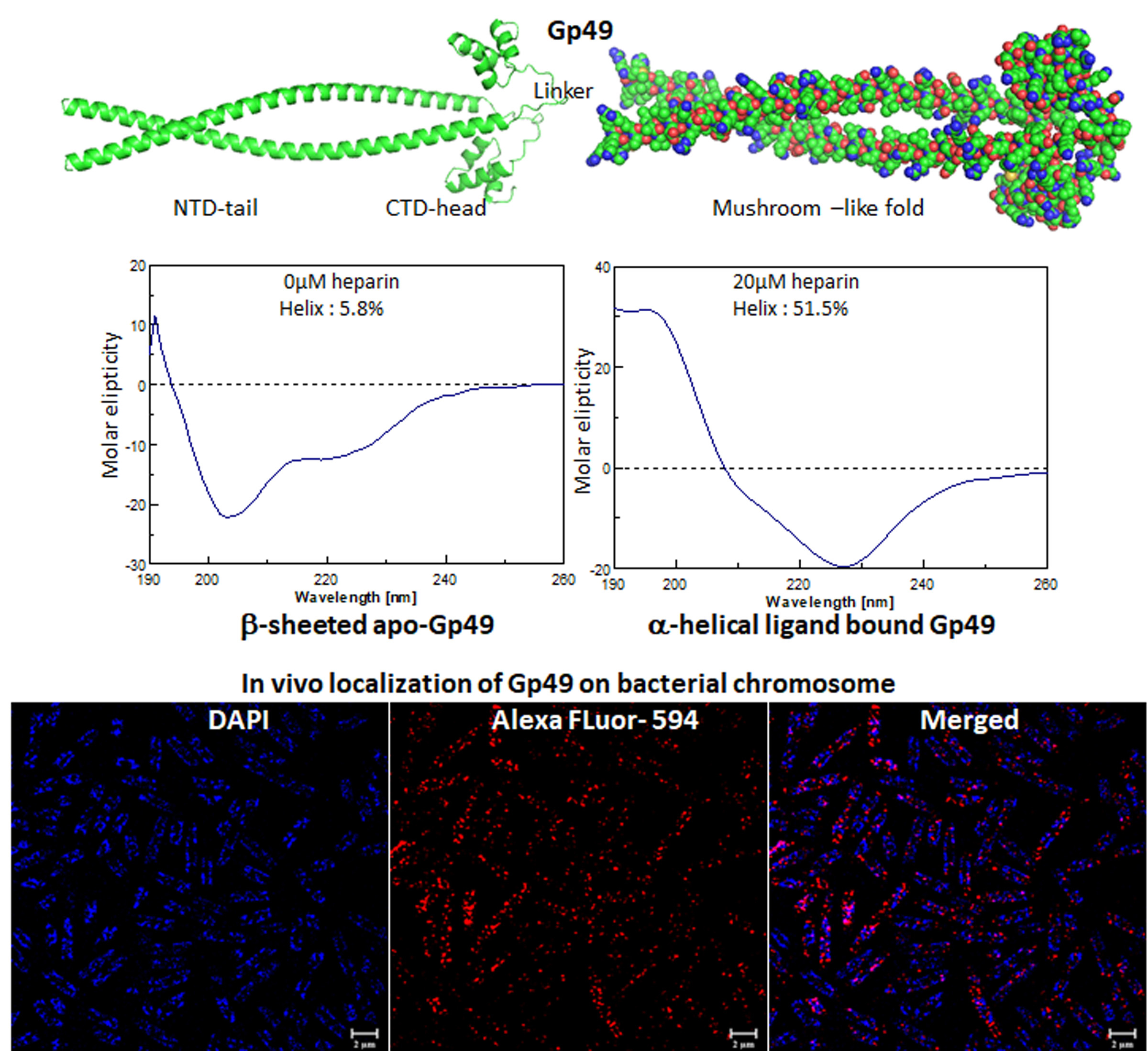
A novel nucleic acid-binding protein, Gp49, from mycobacteriophage with mycobactericidal activity has the potential to be a therapeutic agent (IJBM, 2023).
The mycobacteriophages encode unique proteins that are potent therapeutic agents. We screened several clones with mycobactericidal properties from a genomic library of mycobacteriophages. Here we report the properties of one such clone coding the gene product, Gp49, of the phage Che12. Gp49 is a 16 kD dimeric protein having an HTH motif at its C-terminal and is highly conserved among mycobacteriophages and likely to be part of phage DNA replication machinery. Alphafold predicts it to be an -helical protein. However, its CD spectrum showed it to be predominantly -sheeted. It is a high-affinity heparin-binding protein that has similarities with the macrophage protein Azurocidin. Its -sheeted apo-structure gets transformed into -helix upon binding to heparin. It binds to linear dsDNA as well as ssDNA and RNA cooperatively in a sequence non-specific manner. This DNA binding property enables it to inhibit both in vitro and in vivo transcription. The c-terminal HTH motif is responsible for binding to both heparin and nucleic acids. Its in vivo localization on DNA could cause displacements of many DNA-binding proteins from the bacterial chromosome. We surmised that the bactericidal activity of Gp49 arises from its non-specific DNA binding leading to the inhibition of many host-DNA-dependent processes. Its heparin-binding ability could have therapeutic/diagnostic usages in bacterial sepsis treatment.
.
|
|
In vivo regulation of bacterial Rho-dependent transcription termination by the nascent RNA (JBC 2022).
Bacterial Rho is an RNA-dependent ATPase that functions in the termination of DNA transcription. However, the in vivo nature of the bacterial Rho-dependent terminators, as well as the mechanism of the Rho-dependent termination process, are not fully understood. Here, we measured the in vivo termination efficiencies of 72 Rho-dependent terminators in E. coli by systematically performing qRT-PCR analyses of cDNA prepared from mid-log phase bacterial cultures. We found that these terminators exhibited a wide range of efficiencies, and many behaved differently in vivo compared to the predicted or experimentally determined efficiencies in vitro. Rho-utilization sites (rut sites) present in the RNA terminator sequences are characterized by the presence of C-rich/G-poor sequences, or C>G bubbles. We found that weaker terminators exhibited a robust correlation with the properties (size, length, density, etc.) of these C>G bubbles of their respective rut sites, while stronger terminators lack this correlation, suggesting a limited role of rut sequences in controlling in vivo termination efficiencies. We also found that in vivo termination efficiencies are dependent on the rates of ATP hydrolysis as well as Rho-translocation on the nascent RNA. We demonstrate that weaker terminators, in addition to having rut sites with diminished C>G bubble sizes, are dependent on the Rho-auxiliary factor, NusG, in vivo. From these results, we concluded that in vivo Rho-dependent termination follows a nascent RNA-dependent pathway, where Rho-translocation along the RNA is essential and rut sequences may recruit Rho in vivo, but Rho-rut binding strengths do not regulate termination efficiencies.
|
|
Projects in progress:
- Understanding the physiological consequences of Rho-dependent termination.
- Understanding the role of the omega subunit of RNAP in Rho-dependent termination.
- In vivo localization of Rho and NusG by super-resolution microscopy.
- Identification Rho-RNAP functional interaction domains.
- Isolation and characterization of anti-mycobacterial proteins from mycobacteriophages.
- Design of antiterminator peptides from Psu protein.
Extramural Funding:
- SERB JC Fellowship (2022-2027)
- SERB grant (2024-2027)
- DBT Grant (2022-2025)
Awards/Recognition:
- 2002-2007: GRIP research grant award from NIH, USA.
- 2003-2008: Wellcome Trust, UK, Senior Research Fellowship.
- 2007: DBT Bioscience carrier development award.
- 2007: Elected member of GRC.
- 2008: DST Swarnajayanti Research Fellowship.
- 2011: Elected fellow of NASI, Allahabad.
- 2015: Member DST- SERB, task force.
- 2018: Elected Fellow INSA, New Delhi.
- 2018: Elected fellow IASc, Bangalore.
- 2018: Fellow of Telangana Academy of Sciences.
- 2019: DBT TATA Innovation Fellowship.
- 2021: Elected Fellow of West Bengal Academy of Sciences.
- 2022: SERB JC Bose Fellowship.
Editorial Board Member:
- Journal of Applied Genetics (microbial Genetics section).
Reviewer of Journals/grants/Thesis:
-
Nature Communications, MBio, TIBS, Journal of Bacteriology, Journal of Molecular Biology, Microbiology, PLOS one, Biochemical transactions, Communication Biology, msphere, MEEGID, Microbial Genomics, Scientific Reports, Indian Journal of Biophysics and Biochemistry, Journal of Bioscience, etc.
- The reviewer of grants for different granting agencies like DBT, DST, DRDO, Indo-French programs, Marsden Fund, New Zealand. etc.
- The reviewer of the thesis of the Ph.D. students from renowned institutions like SINP, Kolkata; Bose Institute, Kolkata; IISc., Bangalore; IMTech, Chandigarh, HCU, JNU, etc.
Patents :
- 'NOVEL SYNTHETIC PEPTIDES'; Indian Patent Application No. 201841048582 filed on December 20, 2019.
| |
|
|
|
| Last updated on : Friday, 29th April, 2022. |
|
|
|
|